Integrative analysis of multi-omics data for discovery and functional studies
Dr. Collins’ lab specializes in the generation, processing, statistical analysis, integration, and visualization of high-density genomics and proteomics data for translational research.
Collaborative model
Dr. Collins’ lab expertise can be accessed through the Vancouver Prostate Centre Genomics core facility. The core facility offers expertise to academic and clinical research projects requiring a pan-omics approach. This expertise includes DN and RNA sequencing, proteomics, analytical pharmacology, biostatistics, bioinformatics and computer science. The core has a demonstrated ability to innovate, a competence to manage research projects from experimental design to data interpretation and support the researchers’ grant applications and publications. The core’s highly trained personnel closely collaborate with the users to offer essential recommendations, contributing significantly to the success of the projects. A driving force behind the core’s expanding repertoire of capabilities is its capacity to adapt its expertise to the changing needs of the research projects for a more personalized boutique experience. The competitive service prices are based on a cost-recovery, not-for-profit model to provide economical and personalized solutions.
Benefits of the core facility to research projects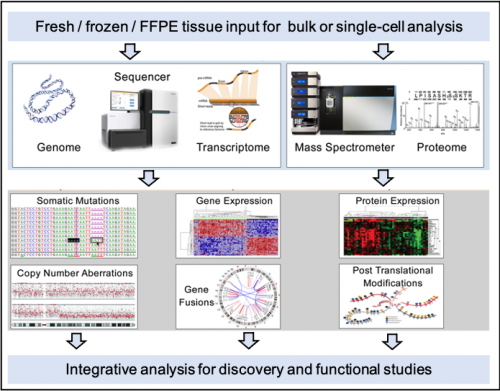
- Researchers capitalize on analytical methods without first becoming experts
- Personalized approach to projects for high-quality results and publication
- Extensive experience and highly trained personnel
- Ongoing internal and external collaborations allow the core facility to improve its expertise continually
- Cost-effective services based on a cost-recovery, not-for-profit model
- Projects of any size are welcome
Laboratory expertise includes
- Experimental design
- Sample preparation (e.g. DNA/RNA extraction and quality control)
- Library preparation and enrichment for bulk DNA and RNA sequencing (does not differentiate among cell types within the sample)
- Single-cell RNA sequencing (10x Genomics)
- Protein quantification by liquid chromatography-mass spectrometry
- Analytical pharmacology
- Pilot projects to evaluate technology performance
- Training and use of instruments
- Use of discrete algorithms
- Statistical analysis
- The computational workflows to perform single-omic and integrated analyses of genomic, transcriptomic and proteomic data.
- High-throughput data analysis is done on a high-performance cluster system.
Currently supported applications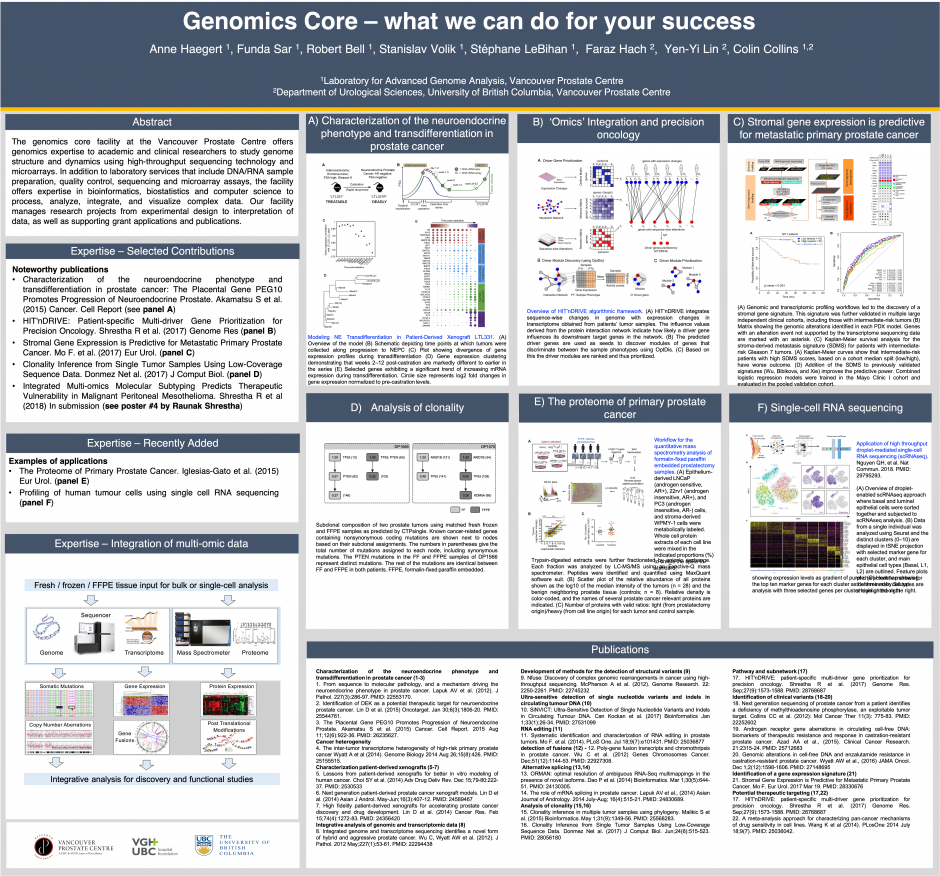
- 10x Genomics single-cell sequencing
- Exome sequencing for integrative analysis of SNVs and CNVs
- RNA sequencing for gene expression or whole transcriptome analysis
- Targeted DNA and RNA sequencing
- Small RNA sequencing
- NGS library sequencing
- Qualitative (label-free) and quantitative (TMT, SILAC) differential analysis of protein levels
- Analytical chemistry and clinical pharmacology
Major equipment
- Maxwell RSC for automated DNA and RNA extraction system
- Agilent TapeStation 4200 for quality control of DNA and RNA
- Covaris M220 focused-ultrasonicator for sample preparation
- ThermoFisher Zephyr NGS liquid handler systems for automated preparation of NGS sequencing libraries
- Illumina NextSeq, MiSeq, MiniSeq
- MGI G50R
- nanoLC Orbitrap Lumos, standard LC triple quad, standard LC with photometric detection for LC-MSMS
- Computing infrastructure: 28 nodes / 896 cores / 2Pb of storage
Contact
Dr. Stéphane LeBihan, PhD
Manager | Genomics core facility | Sequencing, Proteomics and Bioinformatics
Colin Collins lab | Vancouver Prostate Centre
2660 Oak Street, Vancouver BC V6H 3Z6 Canada
Email 1: slebihan (at) mail.ubc.ca
Email 2: slebihan (at) prostatecentre.com
Mobile: 778-991-4506